

Graphs are a great example of domain specific programming systems. There is a lot of underlying structure about graphs and a lot of assumptions can be made about the nature of the data. Since there are so many pre-existing graph search/coloring/building algorithms, a lot of system-level optimizations can be made on a graph as well as algorithmic optimizations. If the user knew nothing of the underlying structure, these optimizations (such as the fact that there are only vertices and edges as datatypes or something of that nature) would not be able to be exploited.
This comment was marked helpful 0 times.

@AnnabelleBlue... one comment I'd like to make is that it's the system that has knowledge of the underlying structure, and that's why the system is able to make sophisticated optimizations. The application programmer/user need not know about about the underlying representation of the graph used by the system. In class we talked about how the implementation of the graph abstraction might use different representations based on the HW platform, or operations performed on the graph by the application.
This comment was marked helpful 0 times.


Another application of intensive graph computation is image processing. SIGGRAPH, for example, make heavy use of parallel architectures / frameworks, like CUDA, to improve the performance of image processing.
This comment was marked helpful 0 times.

@TBC, if you want a particularly interesting example, take a look at 'Building Rome in a Day' - http://grail.cs.washington.edu/rome/. This project used graphs build a 3D network representing a city, using images from Flickr.
This comment was marked helpful 0 times.


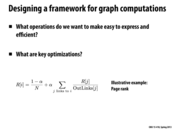
Pagerank is an algorithm that in each iteration, a vertex divides its weight equally to the vertices it links to. We can view pagerank as a case that a person who walks randomly in graph, and the weight of the vertex can be the probability one stay in that vertex. And we always have damping factor \alpha to indicate the probability that one leave the vertex to the next vertex.
This comment was marked helpful 0 times.


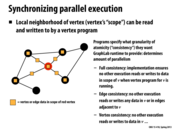
Signalling a vertex is like adding it to the work queue. What this slide is saying is that instead of just signalling all of the vertices once in each iteration (which acts like an implicit barrier), we work on some extreme cases more often by signalling them more than once per iteration, which adds them to the dynamic work queue before the next full iteration over all the vertices.
This comment was marked helpful 0 times.

In this example, when the vertex gets added to the dynamic work queue, there is no specific time when it will get removed from the dynamic work queue. So when the vertex gets removed the worker queue, its neighbors data might not be the latest. Those neighbors might be in the midst of getting their data updated. We would have to do ensure consistency which is not done here if we want to make sure we always have updated data for each and every neighbor.
This comment was marked helpful 0 times.


About graph coloring schedulers: after a graph coloring is found, the vertices are processed one color at a time. Only vertices of the same color are processed in parallel.
This comment was marked helpful 0 times.


The creators of GraphLab use two different types of engines to implement the asynchronous execution model. The first one is called the Chromatic Engine, and it uses vertex coloring to color vertices of a graph so that no two adjacent nodes share the same color. Because nodes of the same color are nonadjacent and can be processed in parallel, the Chromatic fulfills Edge Consistency (see slide 9).
The other engine is called the Locking Engine. Whereas the Chromatic Engine executes some nodes in parallel, the Locking Engine fully executes all nodes asynchronously via a distributed locking system. In particular, this engine utilizes pipelined locking, allowing each machine to request locks for many scopes at once in an effort to hide latency.
For more information about CMU's GraphLab, you can check out the authors' work here.
This comment was marked helpful 0 times.


We need a compare-and-swap in update because it is possible, update(v1,d) and update(v2,d) to be called in parallel in EDGEMAP (the case when two vertices in the frontier, v1 and v2, are adjacent to the same vertex, d, not in the frontier). We use CAS here simply to decide a winner of the race.
This comment was marked helpful 0 times.


On the left, we run F for each edge between the frontier and the unvisited vertices, whereas on the right F only runs once for each unvisited vertex.
This comment was marked helpful 0 times.

In the sparse version, we can see that an vertex will be pulled into the frontier by its in-neighbors, whereas in the dense version, all the vertices will try to put themselves into the frontier. Sparse version is good for sparse graph because there will tend to be less out-neighbors for each vertices in a sparse graph. And therefore compute the new frontier will not use too much time. However, in a dense graph, each vertex may have many out-neighbors, which makes computing the new frontier too expensive.
This comment was marked helpful 0 times.


This requires that EDGEMAP run the function for every edge adjacent to a vertex, not just once. So the early-exit optimization in DENSE_EDGEMAP is unsound.
This comment was marked helpful 0 times.


Green-Marl dose not support operations over arbitrary sets of vertices on each iteration of the traversal, and instead user must explicitly filter out the vertices to skip. Also, the BSF and DFS using in Green-Marl is built-in algorithm whose implementations are built into the compiler.
This comment was marked helpful 0 times.












Domain-specific programming systems also have abundant horse power, but we don't worry about this much because it's difficult to make maximum use of all the available machines. Also, each part of the machine supports different types of parallelism.
This comment was marked helpful 0 times.